Cropping and merging with PDAL
This is the first in a series of posts describing how to use PDAL to develop automated, repeatable processes, the kind that can be tedious to do manually in RiSCAN Pro or other such software. These posts are aimed at a medium-to-high-level user, one who is comfortable with the command line, writing makefiles, and basic scripting. The PDAL tutorials and workshop might be a better start for a beginner user.
You'll need the following softwares for this exercise:
The problem
Given data from five terrestrial laser scanning (TLS) scan positions, crop the data to an area of interest (AOI) and provide those cropped data to the downstream user as a single file.
About as simple as it gets.
The data of interest were collected with a Riegl VZ-6000 on May 01, 2013 in the Tuolumne Meadows area of Yosemite National Park in California. Not bad livin':
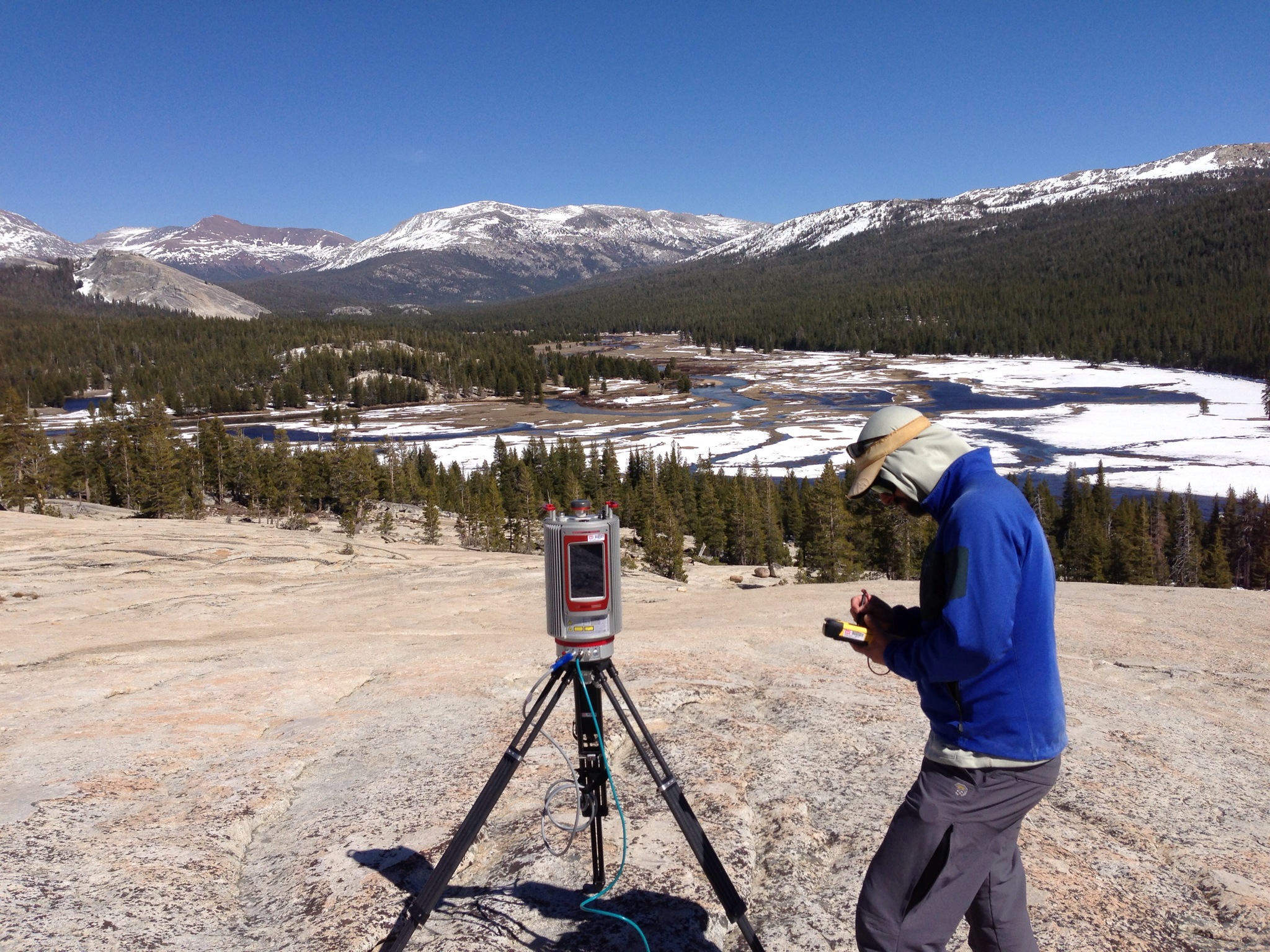

The area of interest is a rectangular box in the east side of the meadow, south of the Lembert Dome:
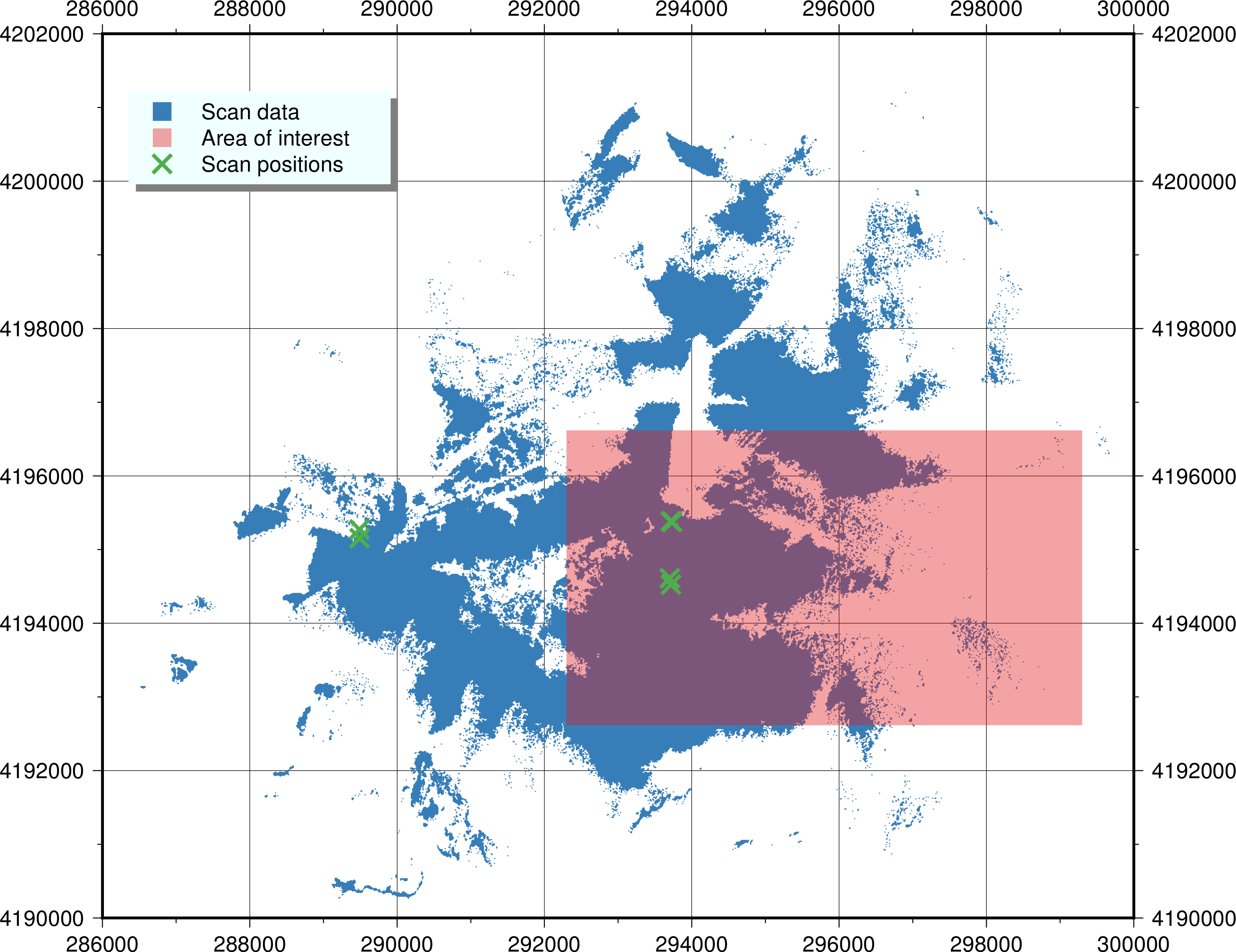
Note: there were actually six scan positions collected that day, but ScanPos006 did not contain any useful data inside the area of interest, so we've not included it in this exercise other than to plot it on the above map.
The process
We pick things up after we've exported a file for each scan position from RiSCAN Pro. We could have used PDAL to do the exporting, but that's a more advanced subject for a later day. I've organized the laz files and our AOI shapefile like this:
$ tree .
.
├── Makefile
├── aoi
│ ├── study_area1.dbf
│ ├── study_area1.gmt
│ ├── study_area1.prj
│ ├── study_area1.shp
│ └── study_area1.shx
└── laz
└── original
├── ScanPos001.laz
├── ScanPos002.laz
├── ScanPos003.laz
├── ScanPos004.laz
└── ScanPos005.laz
We'll use that Makefile to drive our processing.
Cropping
We're going to crop the data with PDAL's crop filter by providing it a WKT polygon of our AOI.
Because our AOI is a shapefile, we need to convert it to WKT with ogr2ogr.
Since shapefiles can be complicated, there's no straightforward way to convert a shapefile to WKT; in this case, however, we know that our shapefile is simple enough to use this simple Python script, located in ~/bin/ogr2wkt (which is on my $PATH):
# !/usr/bin/env python
import sys
from osgeo import ogr
if len(sys.argv) != 2:
print "ERROR: invalid number of arguments"
sys.exit(1)
data = ogr.Open(sys.argv[1])
for feature in data.GetLayer():
print feature.GetGeometryRef().ExportToWkt()
We glue everything together with a couple of make rules (in Makefile):
CROPPED:=$(patsubst laz/original/%.laz,las/cropped/%.laz,$(wildcard laz/original/*.laz))
SRS:=EPSG:32611
cropped: $(CROPPED)
.PHONY: cropped
laz/cropped/%.laz: laz/original/%.laz aoi/study_area1.shp
pdal translate -i $< -o $@ --readers.las.spatialreference=$(SRS) -f crop --filters.crop.polygon=$(shell ogr2wkt $(word 2,$^))
RiSCAN Pro doesn't export laz data with spatial reference information (at least, it doesn't without diving deep into their GeoSysManager, which I've never found worth it).
So we manually specify the source spatial reference system with --readers.las.spatialreference.
We can use make's parallelization to process all the files at once:
make -j 5 cropped
Boom.
We've got five cropped laz files in las/cropped, taking advantage of multiple cores.
Merging
Merging these cropped files into one big deliverable is even simpler with pdal merge.
I still like to use a make rule, for repeatability:
laz/2013-05-01-StudyArea1.laz: $(CROPPED)
pdal merge $^ $@
Conclusion
Our tree should now look like this:
$ tree .
.
├── Makefile
├── aoi
│ ├── study_area1.dbf
│ ├── study_area1.gmt
│ ├── study_area1.prj
│ ├── study_area1.shp
│ └── study_area1.shx
└── laz
├── 2013-05-01-StudyArea1.laz
├── cropped
│ ├── ScanPos001.laz
│ ├── ScanPos002.laz
│ ├── ScanPos003.laz
│ ├── ScanPos004.laz
│ └── ScanPos005.laz
└── original
├── ScanPos001.laz
├── ScanPos002.laz
├── ScanPos003.laz
├── ScanPos004.laz
└── ScanPos005.laz
While it's not hard to do this sort of work in a GUI, things get harder when you want to do the same process over and over, tweaking the inputs, or you want to do the work on a large number of files. Automation such as this is also handy when integrating with other processing steps, such as map generation.